Introducing OpenAPI
11 February, 2015
The OpenAPI project attempts to make it easier to connect people with data. This document discusses some of the challenges faced in connecting people with data, and how OpenAPI addresses these challenges. It describes the technical specifications of the OpenAPI architecture, and provides some examples of OpenAPI in action. This document also discusses projects similar to OpenAPI, and how OpenAPI is distinct from these.
What is OpenAPI: blue sky version
The problem: connecting people with data
OpenAPI's main aim is to make it easier for people to connect with data. Connecting with data is not just a simple case of making raw data available to people (though this is one component of it). When people are fully connected with data they can examine, manipulate, display, compare, interpret and share it. We believe the following things are required to connect people with data:
- Access to data
- Domain knowledge
- Data Science skills
- Statistical Graphics skills
- Graphical Design skills
While many people possess one or several of these attributes, it is very rare to find someone who possesses all of them. Access to data is getting easier thanks to initiatives to make data widely and freely available, like Open Data, Open Government and Open Access. Domain knowledge, data science skills, statistical graphics skills and graphical design skills can all be acquired through education and experience. OpenAPI intends to help connect with data those people who do not have the luxury of acquiring special skills and knowledge. OpenAPI will do this by allowing anyone to contribute using those skills that they already possess.
Possible solutions
One solution to the problem of connecting people with data is providing software or resources containing all of the required attributes on the user's behalf. Some examples of this are:
Public visualisation services
There are a variety of services which present data in visual form for non-experts to use. Some examples of this:
-
Wiki New Zealand captures freely available data about New Zealand and uses it to produce graphical charts. The website producers use their time and skills to represent data in a uniform style. Wiki New Zealand also welcomes contributions from its users, and provides a uniform way of producing graphical output.
Wiki New Zealand allows people to examine, share and interpret data. It does not provide tools for further manipulation of that data, though it does provide data sources, as well as cleaned data in downloadable Excel spreadsheets. Though contributions are welcomed, a contributor will still need to have many of the the attributes listed above to do so. A user who wants to investigate some data who does not have the necessary skills will have to wait until someone else does the work.
-
Gapminder captures data from a variety of sources, including U.N. organisations, government and research organisations, and commercial sources. The creators have developed software tools for creating visualisations of the data, including interactive graphs and videos of data over time. Gapminder invites users to contribute to its project in various ways, including programming and visualisation work. Users are required to register their interest in contributing.
Gapminder enables people to examine, share and interpret data. It does not provide the software tools used to produce its output; instead it suggests that people use Google's Motion Chart and Public Data Explorer tools. Gapminder's data sources are made available, along with links to original sources where data has been cleaned. Data sources are provided for download as Excel Spreadsheets, and for viewing online in Google-hosted spreadsheets. Contributors are asked to list their visualisation and programming skills when registering their interest, meaning that users without the required skills would need to wait for someone else to do the work.
-
Visual programming
Two significant obstacles to connecting people with data are data science skills and statistical graphics skills. Visual programming software attempts to overcome these obstacles by providing graphical interfaces to common data analysis jobs. This way a user can do an analysis by choosing different options from the interface without having to work with any scripts or code.
Visual programming makes massive demands on its authors to ensure it covers all the uses that may be required, and to make sure that the entire program works well. If some piece of analysis or some aspect of an analysis is not coded into the visual programming software it is no small task for a user to have it included.
Examples:
-
Data mining and machine learning software with a visual programming interface written in Python.
Red-R - archived at https://web.archive.org/web/20130620231300/http://red-r.org/
Visual programming interface for the R statistical environment.
-
Comprehensive solutions for advanced users
We recognise that there appears to be some similarities between the OpenAPI system and some of the advanced comprehensive solutions that already exist. One such example is the Galaxy platform for bio-medical research. A discussion of how these systems differ from OpenAPI in both their intentions and their execution follows our description of the details of the OpenAPI system.
The OpenAPI solution: everyone contributes a small amount
Rather than attempt to capture all of the necessary attributes in one program or service, OpenAPI attempts to capture small contributions which can be combined to suit the user. This means that contributors only require some of the attributes that we have listed. This allows for small contributions like:
A contributor with raw data can make this data available to other OpenAPI users.
A domain expert can clean up data and annotate it.
Someone with data analysis skills can produce meaningful statistics about the data.
A statistical graphics guru can provide a script to make suitable charts for a data type.
The graphical design team at a news blog can take graphical output and manipulate it to produce well designed images for their articles.
In OpenAPI each of these contributions is meaningful. Further, they allow for someone who has none of the required attributes to combine other users' contributions in a meaningful way. Even if a contributor has no particular facility in any of the areas mentioned so far, she can still contribute my making data and scripts available to other users of OpenAPI.
At its simplest OpenAPI is about modules. Each module can ask for inputs, describe some work to be done, and produce some outputs. Modules can be combined in pipelines which plug one module's outputs into another module's inputs. The small contributions listed above could each be captured in a module, and a further contribution would combine them as a pipeline.
What is OpenAPI: technical version
The OpenAPI system is comprised of three main parts:
Modules
A module is the smallest piece in the OpenAPI system. It describes a job to be done. This can be range from something simple like passing a data file into OpenAPI, to a complicated script in a language like R or python. Each module has zero or more inputs, and zero or more outputs. Modules are specified using XML.
Pipelines
A pipeline describes how modules' inputs and outputs are connected to each other to perform a series of tasks. A pipeline contains a list of components (modules, or other pipelines) and a list of pipes which describe how to connect component outputs and inputs. Pipelines are specified using XML.
Glue systems
An OpenAPI glue system can read module and pipeline XML and execute the code in modules to produce the desired outputs.
The following sections expand on each of these parts of the OpenAPI architecture in more detail.
Modules
A module is the most basic component of OpenAPI, the smallest piece of work which can be run to produce an outcome. In general a module will describe some code to be executed. The code is executed, not by the OpenAPI glue system, but by some specified platform or service. A module can require zero or more inputs for its source scripts to be executed, and can produce zero or more outputs.
A module is described in XML, with a root element called 'module'. A basic outline of module XML follows:
<module>
<description> ... </description>
<platform name="..." />
<input name="..." type="...">
<format>...</format>
</input>
<source>
...
</source>
<output name="..." type="...">
<format>...</format>
</output>
</module>
<description>It is good practice to include a plaintext description of what a module does with every module. This might explain in plain language what inputs the module requires, what job the module will do, and what outputs the module produces.
Example:
<description> This module summarises an R data frame. It requires an R data frame as input, and produces a PNG image file called "pie-summary.png" <description><platform>Every module requires a platform. For example, a module may wrap an R script, which requires the "R" platform. Similarly a module wrapping a python script would require the "python" platform.
The platform element requires only one attribute to be set:
name: the name of the required platform
Example:
<platform name="python"/><input>A module can contain zero or more named inputs. Inputs can be one of two types: 'internal' inputs are used when the input is an object internal to the module's platform; 'external' inputs are used when the input comes from an external object, e.g. a file on the local file system or on a web server. Inputs require the following attributes:
name: a unique character stringtype: 'internal' or 'external'
Each input also contains a
<format>element, which contains a 'formatType' attribute (this defaults to "text"). The contents of 'format' describe the data type of the input, e.g. R data frame, CSV file.Internal example:
<input name="messyData" type="internal"> <format formatType="text">R data frame</format> </input>External example:
<input name="messyCSV" type="external"> <format formatType="text">CSV file</format> </input><source>A module's sources contain the scripts to be executed. For example, if a module wraps an R script which takes a data frame and produces a cleaned data frame, a source element points to this R script.
A source element can either contain the required script as its content, or provide a reference to the script's location.
A script file's location is provided using the 'ref' and 'path' attributes. 'ref' can provide a resolvable URI to the script, e.g. an absolute or relative file path. If 'ref' provides only a filename, 'path' can be used to define where this filename should be searched for.
A module can have have more than one source element. The order in which these are to be run can be set using the 'order' attribute, which requires an integer value. The sources will be run according to the 'order' value from lowest to highest, with sources with no 'value' defined as having value greater than zero and less than one.
Inline source example:
<source order="0"> <![CDATA[ library(Rgraphviz) Ragraph <- agopen(myGraph, "myGraph") ]]> </source>Referenced source examples:
<source ref="~/Rscripts/computeStatistics.R"/> <source ref="sensibleScriptName.py" path="http://website.url/scripts/python"/><output>Each module can provide zero or more named outputs. These outputs provide the results of executing the module's scripts. The output type can be 'internal' if an object internal to the module's platform is to be passed to another module, or 'external' if the script produces a file.
The following attributes are required for an output:
name: a unique character stringtype: 'internal' or 'external'
and, iftypeis 'internal':ref: the filename of the output
Each output also contains a
<format>element, which contains a 'formatType' attribute (this defaults to "text"). The contents of 'format' describe the data type of the input, e.g. "R data frame", "CSV file".
Pipelines
A pipeline describes how a collection of modules are to be connected together to create more complex jobs. The simplest pipeline describes how the output from one module is to be connected to the input of another module. When the pipeline is run, the first module's source scripts are executed. Its output is then provided to the second module, and the second module's scripts can be executed.
Pipelines can connect a module to a module, connect a module to another pipeline, or even connect two pipelines. Both modules and pipelines are treated as <component>s, which are be connected by <pipe>s.
Each pipe describes the connection of an output from one component to an input of another component. Pipelines can be made increasingly complex by adding more components and connecting them with pipes. Pipelines can be made as complex as the user desires, as long as the various components have all they need to run, and there are no loops in the order of operation.
A pipeline is described in XML, with a root element called 'pipeline'. A basic outline of pipeline XML follows:
<pipeline>
<description>...</description>
<component name="..." ref="..." type="..."/>
<component name="..." ref="..." type="..."/>
<pipe>
<start component="..." output="..."/>
<end component="..." input="..."/>
</pipe>
</pipeline><description>It is good practice to include a description of what a pipeline does, including what it requires to run, and what it produces. The description is written in plain language for people to read.
Example:
<description> This pipeline produces a map of which candidate (Seymour and Goldsmith) received more votes at each polling booth where votes were cast for the Epsom electorate in the 2014 General Election. It requires a CSV of polling places data from http://www.elections.govt.nz/. The pipeline produces a PNG image file of the resulting map. </description><component>A pipeline contains one or more components, which can be either modules or pipelines. A component can either be specified as XML code inside the component element, or by providing a reference to an XML document.
Components described as inline XML must have a
nameattribute set, which should be a unique character string. The inline XML must only contain a<module>or<pipeline>element.Components provided by reference require the following attributes:
name: a unique character stringref: absolute or relative path to, or filename of, referenced XML documentpath(optional): provides a search path for locating the specified XML document
type: 'module' or 'pipeline'
Example of inline component:
<component name="customers_csv"> <module> <description> This is a dummy module to provide the file 'customers.csv'. </description> <platform name="R"/> <output name="Ragraph" type="external" ref="~/data/customers.csv"> <format formatType="text">CSV file</format> </output> </module> </component>Example of referenced component:
<component name="calculateScore" ref="module-657.xml" path="~/moduleFarm" type="module"/><pipe>A pipe describes a connection between one component's output and another component's input. A pipe: (a) connects an 'internal' output to an 'internal' input where both components have the same platform; (b) connects an 'external' output to any 'external' input regardless of either component's platform.
Each pipe must contain a
startand anendelement. Thestartelement must have acomponentattribute and anoutputattribute.componentspecifies by name which of the pipeline's components provides the output for this pipe.outputspecifies by name which of this component's outputs is required. Similarly, theendelement must have acomponentattribute and aninputattribute.Example:
<pipe> <start component="layoutGraph" output="myGraph"/> <end component="plotGraph" input="laidoutGraph"/> </pipe>inputs/outputs
As a pipeline can be passed into another pipeline as a component, a pipeline can also have inputs and outputs. Rather than specifying inputs and outputs in the pipeline's XML these inputs and outputs are determined by the inputs and outputs of its components.
For example, if a pipeline contains a component named 'plotPie' which provides an output named 'twoPies', this pipeline can then provide an output named 'plotPie:twoPies'.
Glue system(s)
A glue system's job is to interpret OpenAPI pipelines and modules, to execute the scripts they provide, and to return the results to the user. There is not a single official OpenAPI glue system; instead we specify here what is expected of a glue system, and provide the details of a prototype glue system.
Requirements
An OpenAPI glue system must do the following:
Read pipeline and module XML
A glue system must be able to read and interpret pipeline and module XML as specified in this document. The glue system must also resolve resource locations referenced in pipeline
<component>and module<source>elements, and read and interpret these resources. Resolving these resource locations may involve searching. The glue system must also resolve component searches.Determine a running order for pipeline components
Pipeline
<component>s can be specified in any order, and not necessarily the order in which they will need to be executed.<pipe>s, too, can be specified in any order. A glue system must be able to determine which components produce the outputs required by other components. The glue system must be able to execute the component code in an order that ensures all required inputs will be available when a module is executedExecute the code in module
<source>elementsA glue system must be able to execute the code specified by a module's
<source>elements in the platform specified by the module. If a platform is not supported by a glue system, the glue system should make this known to the user.Resolve the outputs produced by a module
The glue system is responsible for ensuring that module outputs are addressable once the module's
<source>elements have been executed. For 'internal' outputs it is entirely up to the glue system how this is handled. The only requirement is that the glue system makes a module's outputs available to other modules' inputs.As 'external' outputs are produced by module
<source>code directly, the glue system is responsible both for resolving the location of the output, and for ensuring that a resource exists at this location. Where an 'external' output'srefis an absolute resource location this should be trivial for the glue system to check. Where therefis provided as a relative resource location the glue system will need to resolve the output location dynamically at run-time, check that the resource location exists, and make this location available to other modules.An example of how this requirement might be implemented can be found in the section on
conduit.Resolve and make available module inputs
The glue system is responsible for providing resource locations to module inputs. Every module should have its inputs specified by
<pipe>s in the pipeline. The glue system is responsible for making the resource locations of module outputs available to module inputs according to these<pipes>s.Details of how this is implemented in
conduitcan be found in the section onconduit.-
Pipeline
<component>s and module<source>s can be provided by reference to an external resource through therefattribute. A glue system must be able to search for this resource if therefgiven is not enough to resolve the resource's location. If such a search is performed a glue system must notify the user of this.By default, a glue system should (a) recursively search the file directory of the originating module or pipeline file, then (b) recursively search the file directory from which the glue system was invoked.
The
pathattribute can be used to specify additional locations to be searched. A path can either be searched before, after, or instead of the default search locations.If a resource cannot be located a glue system must alert the user.
What else might a glue system do?
This section describes some features of a glue system that are desirable, but not required by the OpenAPI architecture.
A glue system might also offer:
Create modules and pipelines
A glue system could provide a mechanism for creating modules and pipelines. These could be run alongside modules and pipelines described by OpenAPI XML files.
This could take the form of functions in the same language as the glue system, which allow for the scripting and automation of new modules and pipelines. Alternatively, a glue system with a graphical front-end could provide visual programming tools for creating a pipeline.
Export module and pipeline XML
A glue system could provide a way of saving or exporting loaded or created pipelines or modules to file as XML. Entire pipelines could be exported to be shared online. Single components could be saved or exported to be re-used in other pipelines.
Validate pipelines
A glue system can (and probably should) check that a pipe connects an output to an input of the same
<format>. This validation could be as simple as checking that the<format>values match, and failing otherwise.Visualise pipelines
A glue system could produce graphical representations of pipelines. This might be a simple node and edge graph of components and pipes. At the more complex end of things this could be an interactive representation of components, pipes, and their various elements and attributes.
Return outputs
A glue system might return the resource locations of the outputs produced by its components. Depending on the user's requirements this could return just those outputs which are not consumed as inputs by other components, every output produced by every component, just 'external' outputs, or any conceivable combination.
Examples
Modules
<?xml version="1.0"?>
<module xmlns="http://www.openapi.org/2014/">
<description>Lays out a graphNEL graph using the Rgraphviz package</description>
<platform name="R"/>
<input name="myGraph" type="internal">
<format formatType="text">R "graphNEL" object</format>
</input>
<source><![CDATA[library(Rgraphviz)
Ragraph <- agopen(myGraph, "myGraph")]]></source>
<output name="Ragraph" type="internal">
<format formatType="text">R "Ragraph" object</format>
</output>
</module>This module requires the 'R' platform. It has one 'internal' input, "myGraph", an R graphNEL object. The module contains one source element, specified as inline R script. The module produces one 'internal' output, "Ragraph", an R Ragraph object.
Pipelines
<?xml version="1.0"?>
<pipeline xmlns="http://www.openapi.org/2014/">
<description>This pipeline creates a directed graphNEL graph, lays it out using the Rgraphviz package, and then plots the graph as a PNG file using the gridGraphviz package</description>
<component name="plotGraph" ref="plotGraph.xml" type="module"/>
<component name="createGraph" ref="createGraph.xml" type="module"/>
<component name="layoutGraph" ref="layoutGraph.xml" type="module"/>
<pipe>
<start component="createGraph" output="directedGraph"/>
<end component="layoutGraph" input="myGraph"/>
</pipe>
<pipe>
<start component="layoutGraph" output="Ragraph"/>
<end component="plotGraph" input="Ragraph"/>
</pipe>
</pipeline>This pipeline contains three components, named "plotGraph", "createGraph", and "layoutGraph". The contents of the "layoutGraph" component are given by the 'ref' attribute, which contains the relative file location "layoutGraph.xml", taken to be in the same location as the pipeline file itself, as no search 'path' is given. The 'type' of the "layoutGraph" component is a "module". This module is the same as in the example above.
The other two components, also modules, can be examined at simpleGraph/plotGraph.xml and simpleGraph/createGraph.xml.
This pipeline contains two pipes. The first connects the output "directedGraph" in the component "createGraph" to the input "myGraph" in the component "layoutGraph". The second pipe connects the "layoutGraph" output "Ragraph" to the "plotGraph" input "Ragraph".
The following image shows a graphical representation of the pipeline's three components and two connecting pipes:
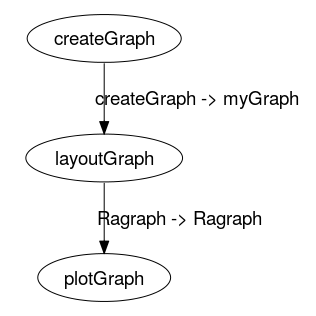
Graph of pipeline example
Glue system: conduit
The conduit package is a prototype built to demonstrate the features of an OpenAPI glue system. It is available as a package for the R statistical environment.
Basic usage: conduit in action
This section shows how the pipeline example above can be read into and executed in conduit.
Installing conduit
The conduit package was built using R version 3.1.x on a 64-bit Linux machine. It requires the following R packages be installed: XML, graph, RBGL, and RCurl. The devtools package is required to install conduit directly from github. The Rgraphviz and gridGraphviz packages are required to run the modules in this example.
Version 0.1 of the conduit, used in this report, is available for download here.
Source code for conduit is available at https://github.com/anhinton/conduit for those who would like to build and install the package manually.
To install the conduit v0.1 using devtools:
devtools::install_github("anhinton/conduit@v0.1")Load the conduit package:
library(conduit)Reading a pipeline from an XML file
The loadPipeline() function is used to read a pipeline XML file into conduit. The loadPipeline() function requires a name argument, and a ref argument which describes the XML file location. The function returns a pipeline object.
We will load the pipeline used in our pipelines example above. The pipeline XML file is in a sub-directory of the R working directory called 'simpleGraph'.
## load the simpleGraph pipeline
simpleGraph <-
loadPipeline(name = "simpleGraph", ref="simpleGraph/pipeline.xml")According to its description:
This pipeline creates a directed graphNEL graph, lays it out using the Rgraphviz package, and then plots the graph as a PNG file using the gridGraphviz packageRun a pipeline in conduit
The runPipeline() function is used to run a pipeline object in R. It requires a pipeline object as its only argument. This function will create a directory for the pipeline in the 'pipelines' sub-directory of the current working directory. If a 'pipelines' sub-directory does not exist it will be created. Each module in the pipeline will create output in a named directory found in ./pipelines/PIPELINE_NAME/modules.
## run the simpleGraph pipeline
runPipeline(simpleGraph)This creates the following files:
- pipelines/simpleGraph/modules/createGraph/directedGraph.rds
- pipelines/simpleGraph/modules/createGraph/script.R
- pipelines/simpleGraph/modules/layoutGraph/Ragraph.rds
- pipelines/simpleGraph/modules/layoutGraph/script.R
- pipelines/simpleGraph/modules/plotGraph/example.png
- pipelines/simpleGraph/modules/plotGraph/script.R
File number 5, pipelines/simpleGraph/modules/plotGraph/example.png is the output file we require, the PNG image of the graph. The image is shown below:
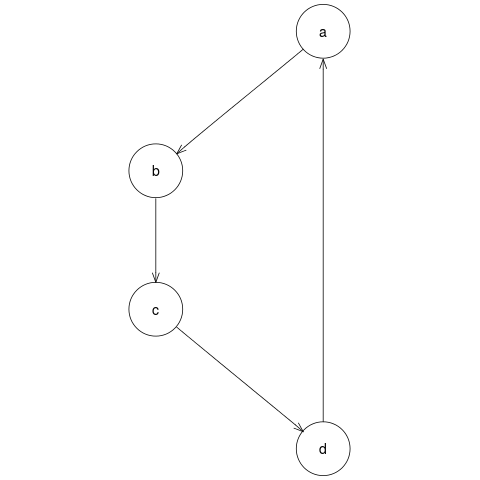
Pipeline output: PNG image file of graph
Technical details
We will further explore the different features of an OpenAPI glue system by examining conduit more closely. An R package, conduit provides functions for reading, running and creating OpenAPI modules and pipelines.
The conduit package can be installed from https://github.com/anhinton/conduit
Read pipeline and module XML
The functions
loadPipeline()andloadModule()read pipeline and module XML from files.Determine a running order for pipeline components
The
conduitpackage'srunPipeline()function determines a running order for a pipeline's components, and then executes the components in this order.Execute the code in module
<source>elementsThe function
runModule()will execute the code specified in a module's<source>elements. Each platform supported byconduithas a platform support function which generates a script file in the appropriate platform language. This script file contains code to import the module's inputs, the code provided by the module's<source>elements to be executed, and code to export the module's outputs. This script is then executed by the platform using R'ssystem()function. Theconduitpackage currently supports the 'R', and 'python' platforms for module<source>code.Resolve the outputs produced by a module
The
conduitpackage uses the local filesystem to store and address outputs produced by modules. When a module is executed usingrunModule()conduitgenerates a directory where the platform support functions will export the module's outputs. 'Internal' outputs are exported as a file format common to the platform. 'External' outputs which are not specified with an absolute locationrefare also saved to this directory.runPipeline()uses this filesystem structure to resolve the addresses of module outputs at run-time.As at 2014-10-29
conduitdoes not support the requirement to ensure that 'external' outputs exist.- Resolve and make available module inputs
runPipeline()determines where all of a pipeline's module inputs will be located based on the outputs produced byrunModule(). As inputs only exist at run-time, inputs for a given module are provided to the module as it is executed. Resolve component searches
When a pipeline
<component>or a module<source>is provided by aref,conduitattempts to resolve the resource location. BothloadPipeline()andloadModule():Treat the reference as an absolute location, and return the resource if it is available at this location.
Otherwise, treat the reference as a location relative to the file location of the XML document which gave the reference, and return the resource if it is available at this location.
Otherwise, treat the reference as a resource name to be searched for, either at the default paths, or at a path determined by the resource's
pathattribute. Theconduitpackage will return the first hit in its search. As at 2014-10-29conduitreturns a warning if the search finds multiple matches, but does not meet the requirement of alerting the user whenever a search is started.
Create modules and pipelines
The
conduitpackage provides the functionsmodule()andpipeline()for creating new modules and pipelines respectively. The resulting objects can be run in the same fashion as modules of pipelines loaded from XML.Export module and pipeline XML
The
conduitpackage provides thesaveModule(),savePipeline()andexportPipeline()functions to save modules and pipelines as XML files.exportPipeline()saves the pipeline and all of its components as XML files in the same directory so the entire pipeline can be shared.Provide a uniform way of executing module code
The
conduitpackage provides therunModule()function as a uniform way of executing module<source>code. The function, rather than the user is responsible for executing the<source>code using the appropriate platform support.
Discussion
How is OpenAPI different?
We are aware that OpenAPI bears some resemblance to a variety of other projects in the broad categories of data access and analysis. It is our opinion, however, that several features make OpenAPI unique.
There are too many other systems to line them all up and knock them all down, so this section provides a comparison between OpenAPI and a selection of related projects that represent a broad range of alternatives.
There are several projects that are structurally similar to OpenAPI's use of modules and pipelines: Mortar + Luigi is a platform for building and running data pipelines; Flow-based Programming is a programming paradigm which treats applications as opaque processes which pass data to other processes through externally defined connections; Web Services Description Language (WSDL) is an XML-based language used to define what is offered by a web service.
Unlike these projects, OpenAPI is intended to be simple; it is aimed at non-technical users, and not at developers. It does not attempt to define data types, does not permit looping, and is not meant to provide a paradigm for program design. Rather than attempting to solve the entire problem of connecting people with data through complex and exhaustive programming, OpenAPI merely specifies a framework for describing working with data. The XML used to describe modules and pipelines is simple enough for a user to type out herself. OpenAPI is able to combine these simple pieces to provide as much complexity and power as its users imagine but it is aimed primarily at small scale, single person projects rather than large scale production systems.
Some projects offer hosted web services and collaboration tools for data analysis: OpenCPU provides an HTTP API for performing data analysis using R; RCloud is a web-based platform intended for producing interactive web-based graphics; ElasticR provides a platform for cloud-based data analysis. Other projects offer bundled, comprehensive data workflow solutions: KNIME offers a graphical environment for assembling pipelines of data analysis modules for pharmaceutical and life science research; VisTrails is a workflow management system for scientific data exploration and visualisation; Alteryx provides a graphical environment for data manipulation and analysis for business users.
OpenAPI is not an attempt to provide a single, comprehensive solution to the problem of connecting people with data, nor is it an attempt to combine existing solutions in domain-specific contexts. Instead OpenAPI allows users to combine the contributions of other users, and to share these combinations in turn and build small contributions. OpenAPI is unique in that it is possible to make small but meaningful contributions. A contributor can do something as simple as writing an XML module for a piece of code, and will have made a meaningful contribution. Another contributor could write a pipeline which connects the contributions of several modules from various sources. Yet another contributor could write a small script and that, even without intending to be, is a contribution in the OpenAPI model.
The Galaxy Project also attempts to connect people with data, specifically in the field of biomedical research, and offers both a hosted and an installable platform for this. Galaxy is a web-based platform. This means most users will need nothing more than their regular web browser to use Galaxy. Galaxy solves the problem of access to data by hosting data sets, and then providing the option to share these data sets on public servers. The problems of domain knowledge, data science skills, and various graphical skills are not solved directly by Galaxy. Rather, these are addressed through Galaxy users sharing their data, workflows, and visualisations through the Galaxy Pages service. These Pages allow other users to examine and reproduce the workflows of other users. This then serves as both a published record of the findings of a project, and an educational tool for helping users create work of their own.
Galaxy differs from OpenAPI by being specifically aimed at biomedical research. Like OpenAPI, a user can make use of someone else's workflow to pursue her own research. However, a high level of domain knowledge and skills are still required to make use of and understand the platform. Unlike OpenAPI pipelines, Galaxy workflows must be run on Galaxy servers, using the tools available on this platform. OpenAPI pipelines can be run in any glue system, and can make use of any platforms that the glue system supports. OpenAPI glue systems can be anything from a self-contained application on a single machine, to a distributed system of platform resources over a local or worldwide network.
Ubuntu Juju uses a visual environment to connect applications and services to each other by representing them as nodes called charms, and connecting these nodes. While the metaphors are similar, Juju is aimed at large-scale pipelines, abstracting across whole systems; OpenAPI is meant for much smaller pipelines.
Finally, OpenAPI does not insist on there being one solution, one glue system, one server on which all the work must take place. Rather it allows for modules and pipelines to be brought into already existing setups. Developers are encouraged to provide support for additional platforms in existing glue systems, or even to create whole new glue systems with whatever features they desire. Because module and pipeline XML wraps around existing scripts a contributor's work is not trapped in OpenAPI; the scripts remain in plaintext to be copied and pasted anywhere one pleases. Contributors do not have to learn whole new languages and methods of programming to begin participating in the OpenAPI project, nor do their solutions become locked into a particular service or product.
The unintended benefits of OpenAPI
Though not the original aim of the project, we have so far noticed the following benefits to working in modules and pipelines:
Structured data analysis workflow:
Imposing the structure of modules and pipeline on a data analysis task can help keep an analyst's scripting clean and organised. This can help with designing an analysis, as well as making the analysis more readable and understandable.
Visualising data analysis workflow:
OpenAPI pipelines can be visualised as node and edge graphs. This allows us to understand the sequence of tasks in an analysis, and which parts of an analysis depend on other parts.
Challenges and future work
One of the challenges OpenAPI faces is ensuring that a module's system requirements are met. While a module's platform program may be present on the user's system this is no guarantee that the program will meet the requirements of the module's source scripts. In some cases these requirements can be met within the modules scripts; for example, a module on the "R" platform might require a package from CRAN be installed: the module could be set up to first run a script to check if the appropriate package is installed, and could even install it.
Other module scripts might make greater demands on a system, and may require specific versions of programs and their dependencies be installed. One solution to this might be to provide a mechanism whereby a module's sources can be executed by the module's platform program on another machine. An IP address and authentication credentials (or similar) could be specified along with the module's platform, and the addressed machine could be set up so as to guarantee the module script in question can be executed. This has the drawback of the machine needing to be instantly available to the user.
Two programs for creating virtual machine environments, Docker and Vagrant, might provide machines which meets the requirements of a module. Both of these programs are capable of providing virtual machines on a user's machine, and both can be set up via provisioning scripts to provide uniform environments to meet a module's requirements. Both also provide networks interfaces which could be addressed in the way speculated for a remote machine.
Furthermore, OpenAPI glue systems themselves may have certain system requirements. It is conceivable that an entire glue system might be run on a remote machine or in a virtual environment like Docker or Vagrant.
Further reading/links
A guide to using
conduitwith detailed descriptions and examples for each function is available. The guide also contains a fully worked example of how to create, execute, and share an OpenAPI pipeline usingconduit.
https://anhinton.github.io/usingConduit/usingConduit.htmlResources from Paul Murrell's talk, "The OpenAPI project", given in the Department of Statistics, University of Auckland, in 2014.
https://www.stat.auckland.ac.nz/~paul/Talks/OpenAPI2014/- The example modules and and the 'simpleGraph' pipeline used in this document:
Summary
In attempting to connect people with data, OpenAPI addresses the need for domain knowledge, and skills in data science, statistical graphics, and graphic design. The OpenAPI solution is to allow meaningful contributions from people who have some of these skills, rather than requiring that a user have all of them. In allowing small contributions OpenAPI invites more people to be involved. Those without any particular data skills can make use of the contributions and examples of those who do.
The OpenAPI architecture consists of modules, pipelines, and glue systems. A module is the smallest piece of OpenAPI, meant to encapsulate a simple data task. Pipelines are arrangements of modules, linked together to produce more complex outcomes. Glue systems are software which can interpret modules and pipelines, and can execute the scripts within.
Ashley Noel Hinton was supported by a Univeristy of Auckland Faculty of Science Research Development Grant.
This document was produced using the knitr package for R, and the pandoc document conversion program. The document source is available for download.

Introducing OpenAPI by Ashley Noel Hinton and Paul Murrell is licensed under a Creative Commons Attribution 4.0 International License.
References
“About Gapminder.” Gapminder. http://www.gapminder.org/about-gapminder/.
“Alteryx.” Alteryx, Inc. http://www.alteryx.com/.
Carey, Vince, Li Long, and R. Gentleman. RBGL: An Interface to the BOOST Graph Library. http://www.bioconductor.org.
Chinnici, Roberto, Jean-Jacques Moreau, Arthur Ryman, and Sanjiva Weerawarana, eds. 2007. “Web Services Description Language (WSDL) Version 2.0 Part 1: Core Language.” W3C Recommendation. W3C. http://www.w3.org/TR/wsdl20.
“Docker.” Docker, Inc. https://www.docker.com/.
“ElasticR.” Cloud Era Ltd. https://www.elasticr.com/welcome.
“Galaxy Wiki.” https://wiki.galaxyproject.org/.
Gentleman, R., Elizabeth Whalen, W. Huber, and S. Falcon. Graph: Graph: A Package to Handle Graph Data Structures.
Hansen, Kasper Daniel, Jeff Gentry, Li Long, Robert Gentleman, Seth Falcon, Florian Hahne, and Deepayan Sarkar. Rgraphviz: Provides Plotting Capabilities for R Graph Objects.
Hinton, Ashley Noel, and Paul Murrell. 2014. Conduit: Prototype Glue System for Openapi.
“KNIME.” Knime.com AG. http://www.knime.org/about.
Lang, Duncan Temple. 2013. XML: Tools for Parsing and Generating XML Within R and S-Plus. http://CRAN.R-project.org/package=XML.
———. 2014. RCurl: General Network (HTTP/FTP/.) Client Interface for R. http://CRAN.R-project.org/package=RCurl.
“Luigi.” Mortar. https://help.mortardata.com/technologies/luigi.
Morrison, J Paul Rodker. “Flow-Based Programming.” http://www.jpaulmorrison.com/fbp/.
Murrell, Paul, and Ashley Noel Hinton. 2014. GridGraphviz: Drawing Graphs with Grid. http://R-Forge.R-project.org/projects/gridgraph/.
“OpenCPU.” https://www.opencpu.org/.
“Orange.” University of Ljubljana. http://orange.biolab.si/.
Python Software Foundation. 2014. The Python Language Reference (version 2.7). https://docs.python.org/2.7/reference/index.html.
R Core Team. 2014. R: A Language and Environment for Statistical Computing. Vienna, Austria: R Foundation for Statistical Computing. http://www.R-project.org/.
“RCloud.” AT&T. http://stats.research.att.com/RCloud/.
“Ubuntu Juju.” Canonical, Ltd. https://juju.ubuntu.com/.
“Vagrant.” HashiCorp. https://www.vagrantup.com/.
“VisTrails.” http://www.vistrails.org/index.php/Main_Page.
Wickham, Hadley, and Winston Chang. 2014. Devtools: Tools to Make Developing R Code Easier. http://CRAN.R-project.org/package=devtools.
Xie, Yihui. 2014. Knitr: A General-Purpose Package for Dynamic Report Generation in R. http://yihui.name/knitr/.